In the intricate realm of molecular biology, understanding the structural dynamics of proteins is paramount for unraveling their functions and devising targeted treatments for diseases. Now, scientists at Brown University have pioneered a groundbreaking technique harnessing the power of machine learning to rapidly predict multiple configurations of proteins, heralding a new era in drug discovery and therapeutic development.
Published in Nature Communications, the study introduces an innovative approach that promises to transform our comprehension of protein dynamics and revolutionize the identification of potential drug targets. Led by Gabriel Monteiro da Silva, a Ph.D. candidate in molecular biology, cell biology, and biochemistry, and Brenda Rubenstein, an associate professor of chemistry and physics, the research team leveraged computational methods and artificial intelligence to probe the intricate choreography of protein molecules.
Traditionally, structural biologists have grappled with the challenge of capturing the dynamic nature of proteins, which undergo shape-shifting transformations crucial for their biological functions. Conventional methods, while adept at depicting static protein structures, falter in capturing the nuanced interplay of proteins in motion—a deficiency that hampers the quest for effective drug targets.
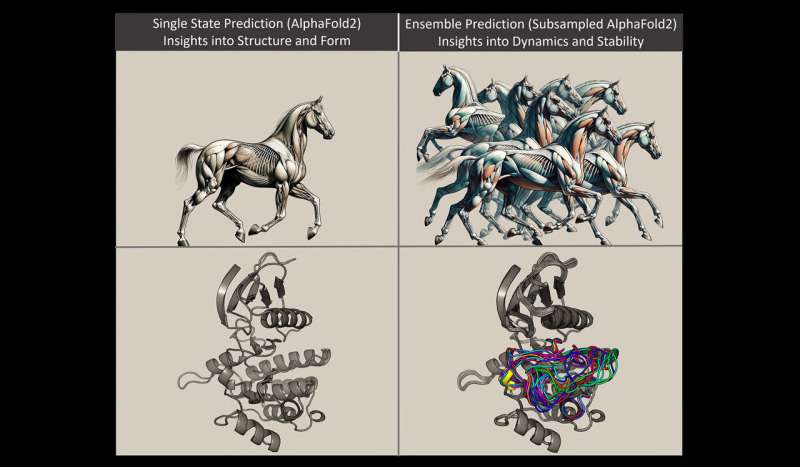
Enter the novel approach pioneered by Monteiro da Silva and his colleagues, which transcends the constraints of static modeling to embrace the fluidity of protein dynamics. Drawing an analogy to a galloping horse, Monteiro da Silva elucidates how proteins, akin to the muscular movements of a horse in motion, adopt diverse conformations dictated by their underlying molecular interactions.
Central to their methodology is the adaptation of AlphaFold 2, an A.I.-powered computational tool renowned for its prowess in protein structure prediction. By manipulating evolutionary signals embedded within protein sequences, the researchers unlocked the ability to swiftly generate multiple snapshots of protein conformations, offering a comprehensive glimpse into their dynamic behavior over time.
This paradigm shift in protein modeling holds profound implications for drug discovery, particularly in the realm of targeted cancer therapy. By elucidating the myriad conformations adopted by key protein targets implicated in disease progression, scientists can unravel the intricate mechanisms underpinning drug efficacy and resistance, paving the way for more tailored and efficacious treatments.
Rubenstein emphasizes the transformative potential of their approach in deciphering the enigmatic interplay between drugs and proteins. By transcending the limitations of static modeling, researchers gain unprecedented insights into how drugs interact with diverse protein conformations, shedding light on the intricate pharmacological mechanisms governing therapeutic outcomes.
Moreover, the newfound efficiency of the proposed methodology promises to accelerate the pace of discovery, circumventing the cost and time constraints associated with traditional computational methods. What once took years of painstaking analysis can now be accomplished in a fraction of the time, empowering scientists to explore the vast landscape of protein dynamics with unparalleled speed and efficiency.
As the research community embarks on this transformative journey into the realm of protein dynamics prediction, the stage is set for a paradigm shift in drug discovery. Armed with cutting-edge computational tools and a deeper understanding of protein behavior, scientists are poised to unlock new frontiers in medicine, offering hope for more effective treatments and improved patient outcomes.
In essence, the dawn of predictive protein dynamics heralds a new era of precision medicine, where targeted therapies tailored to the individual molecular signatures of diseases hold the promise of a brighter, healthier future for all.